Immuno-Oncology Research
Genomic Tools for Characterizing Tumors
Immuno-oncology is an emerging field that has taken great strides in the fight against cancer, bolstered by a refined understanding of how tumors evade the natural immune response. Leading immuno-oncology researchers are leveraging next-generation sequencing (NGS) to study immunotherapy response factors, discover biomarkers, and apply genomics to personalized immunotherapy.
Research into the mechanisms tumors use to evade the immune response have led to promising therapeutic targets. These therapies boost the ability of the immune system to target cancer, or limit the tumor’s ability to evade the immune response. In addition, NGS can help identify which pathways are activated in the tumor environment and involved in processes such as cancer cell proliferation, survival, invasion, and metastasis.
Advancing Cancer Immunotherapy Research
See how the high-throughput capability of NGS enables immuno-oncology biomarker identification.
View InfographicNGS-Based Immuno-Oncology Research Solutions
Studies have shown that response to certain cancer immunotherapies depends on numerous factors. NGS can be used to monitor these dynamic interactions, in real time and with high analytical sensitivity.
Featured Cancer Immunotherapy Research

Developmental Biology Meets Immunotherapy
Listen to Dr. Guo-Liang Chew of the Cancer Science Institute of Singapore discuss a novel role for developmental genes in regulating the immune response to cancer.
Listen to Podcast
Immunotherapies Using Tumor-Specific HLA Ligands
RNA-Seq and HLA typing using NGS are powering the Immatics target discovery platform.
Read Interview
Cancer and Immune System Research Review
High-throughput sequencing has shown remarkable utility in cancer and immunology research, as well as in the development of individualized immunotherapy.
Read ReviewImmune Profiling of Human Tumors
Swetha Anandhan from MD Anderson Cancer Center studies the mechanisms that bolster the ability of malignant cells to evade checkpoint blockade immunotherapy. Her talk highlights the use of single cell RNA-sequencing to identify a unique population of macrophages in glioblastoma multiforme that persists after treatment with immune checkpoint inhibitors.
View Webinar

Advantages of RNA-Seq in Immuno-Oncology Research
Transcriptome analysis is helping to advance cancer immunotherapy research. This white paper demonstrates the performance of Illumina TruSeq RNA Exome compared to a hybridization-based digital counting method (Nanostring Technologies) for quantitative analysis of 57 tumor tissue samples.
Read White PaperNGS Workflows for Immunotherapy Research
Illumina offers several library preparation and sequencing options with access to data analysis options for immuno-oncology research. Streamlined workflows and flexible kit configurations accommodate multiple study designs.
Approximately 90% of the world’s sequencing data are generated using Illumina sequencing by synthesis (SBS) chemistry.*
Click on the below to view products for each workflow step.
Assay targeting multiple variant types, including tumor mutational burden and microsatellite instability, even from low-quality samples.
TruSeq RNA ExomeThe TruSeq RNA Exome library prep kit provides a low-cost solution for analyzing human RNA isolated from limited or low-quality samples, including FFPE.
TruSeq DNA ExomeTruSeq DNA Exome is a cost-effective library preparation and exome enrichment solution.
Targeted RNA expression research panel investigating 395 genes involved in tumor-immune system interactions.
Illumina DNA Prep with EnrichmentOn-bead tagmentation chemistry is combined with a simplified, single hybridization protocol to reduce total workflow time.
TruSeq Stranded Total RNA Library Prep KitA robust, highly scalable whole-transcriptome analysis solution for a variety of species and sample types.
Flexible configurations that support up to 12 exomes per run.
NextSeq 1000 & 2000 SystemsThese cost-efficient, user-friendly, mid-throughput benchtop sequencers offer extreme flexibility to support both current and emerging applications.
Flexibility and high throughput for virtually any genome, sequencing method, and scale of project.
User-friendly data analysis apps for RNA-Seq and other common methods.
DRAGEN Bio-IT PlatformAccurate, ultra-rapid analysis of NGS data, on-premise or in the cloud.
Uses the Isaac Genome Alignment Software and the Isaac Variant Caller for exome data analysis.
Multiomic Sequencing Applications
10X Genomics Chromium Single Cell Multiome ATAC + Gene Expression
Unify single-cell gene expression and chromatin accessibility to help reveal cellular mechanisms driving gene regulation.
Read Tech NoteMeasure Protein Expression Along with RNA-Seq with BioLegend
Learn how to incorporate protein detection into bulk RNA-Seq and develop a workflow for BEN-Seq, providing a holistic approach to cell analysis.
Read Application NoteProbe the immune system at single-cell resolution with 10x Genomics
A multiomic approach to determine how the adaptive immune system functions
Read Tech NoteEnabling single-cell proteomics using TotalSeq-A and NGS with BioLegend
Optimized use of BioLegend TotalSeq antibodies for detection of cell surface proteins and RNA transcripts in cell populations
Read Application NoteInterested in receiving newsletters, case studies, and information on cancer genomics?
Additional Resources

Identifying Immunotherapy Resistance Biomarkers
Eliezer Van Allen, MD highlights how NGS can help identify biomarkers of response and resistance to cancer immunotherapy.

Equipping the Body to Fight Cancer From Within
Recent years have seen an explosion in the translation of immuno-oncology research into new treatment modalities.
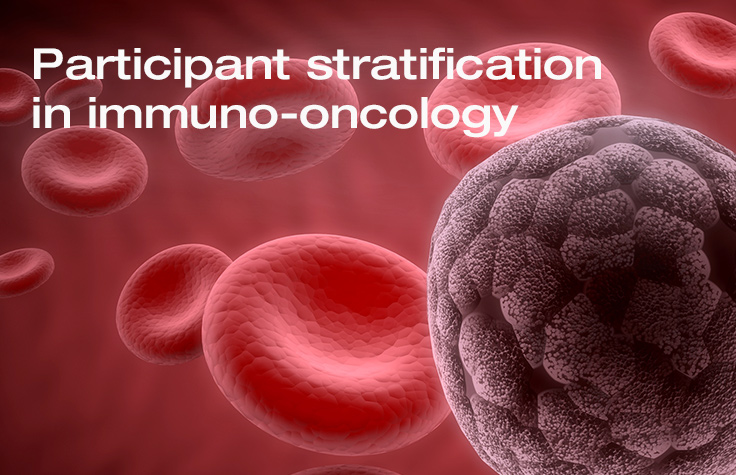
Participant Stratification in Immunotherapy Research
Scientists discuss how advances in immuno-oncology research resulting from NGS help better stratify subjects for clinical trials.

Keeping Pace with Immuno-Oncology Research
NGS technology can accelerate biomarker identification and bring down costs for research subject screening and safety monitoring.

Unmasking the Viral Etiology of Cancer and Immune Disease
Dr. Emilie Hultin and her team use Illumina sequencers to identify novel HPV types correlated with non-melanoma skin cancers.
Using NGS to Aid the Immune System in Targeting Cancers
An overview of exciting new fields of immunotherapy research, and how NGS helps to move them forward.

Immuno-Oncology Research Informatics Applications Using NGS
Improved informatics tools enable neoantigen discovery and tumor microenvironment analysis.

Immune repertoire sequencing on NextSeq 1000/2000 Systems
Learn about immune repertoire sequencing and discover how to obtain high-quality reads with more samples per run in this app note.
*Data calculations on file. Illumina, Inc., 2015