
Epigenetic breakthroughs with the NextSeq 1000/2000 Systems
Learn about innovations of the NextSeq 1000/2000 workflow, including the use of 3D genomics to study the functional context of GWAS variants, in this webinar.
Advance your research
Proven technology to power your discovery
Your email address is never shared with third parties.
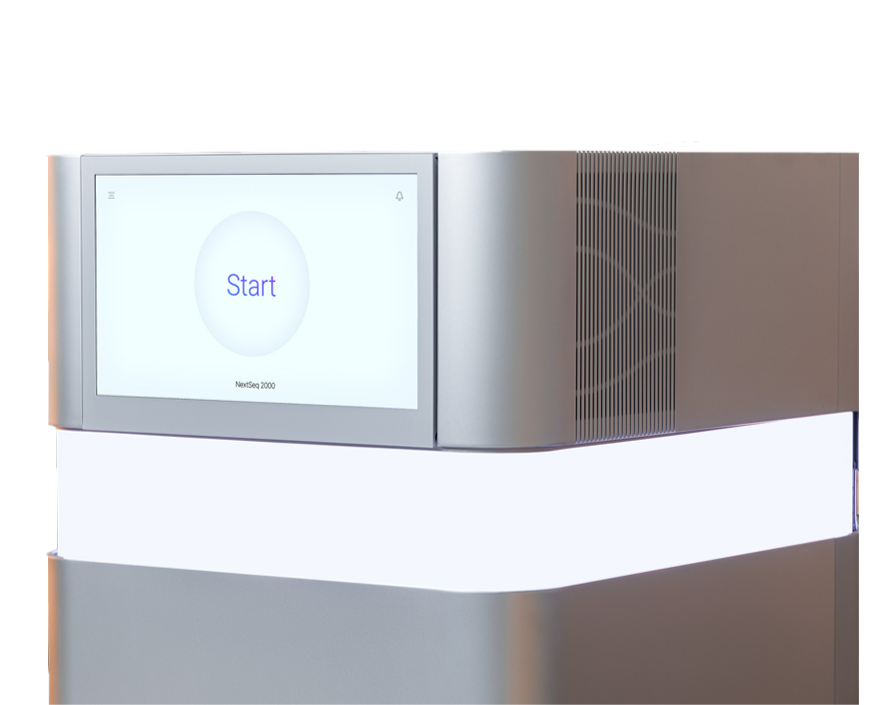
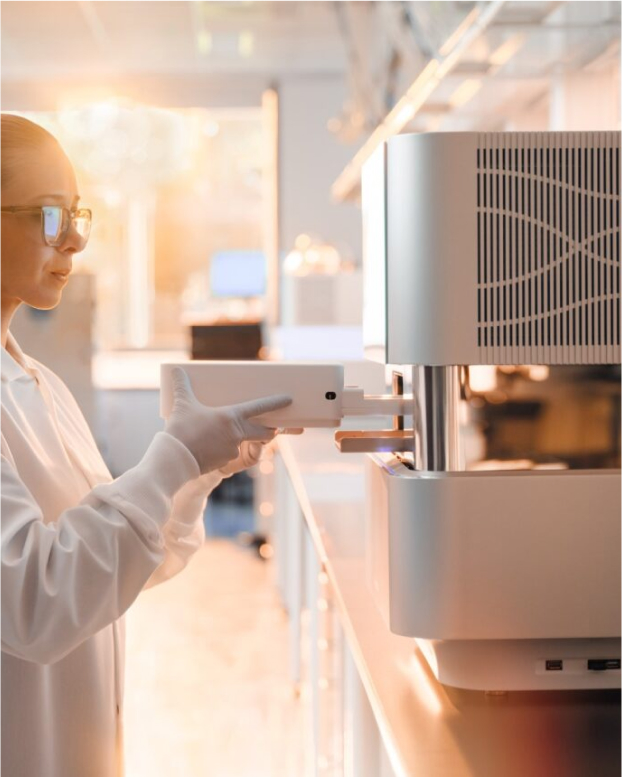


The NextSeq 1000 and NextSeq 2000 Systems are thoughtfully designed to enable more insights on your benchtop. With 14 configurations and read lengths from 1 × 50bp to 2 × 300bp, these systems efficiently handle benchtop workflows while driving down costs and minimizing waste. Confidently expand your research with the NextSeq 1000 and NextSeq 2000 Systems.
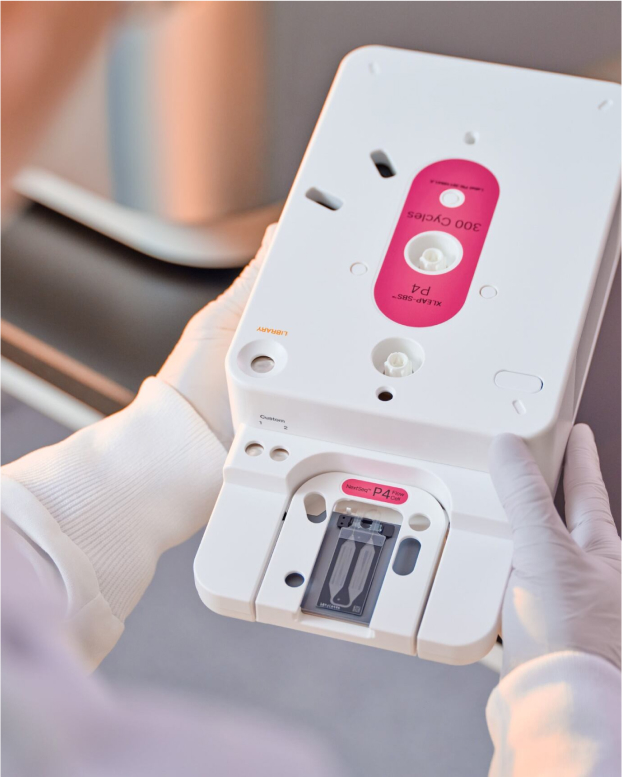

These systems offer operational ease and onboard informatics, simplifying workflows for both new and advanced users. Advanced XLEAP-SBS chemistry, built on the proven foundation of standard Illumina SBS chemistry, enables faster, more economical, and higher quality sequencing than ever before.
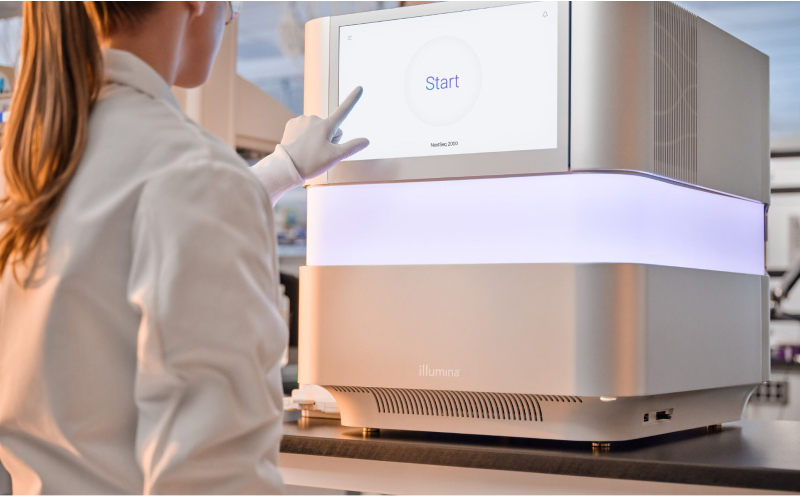

The NextSeq 1000 and NextSeq 2000 Systems offer the certainty of a field-tested solution and a reliable sequencing partner. Take advantage of a large ecosystem of applications, protocols, and informatics built in collaboration with thousands of researchers and thought leaders across the globe.
Access a wide range of applications including single-cell, whole-exome, and RNA sequencing—all on the market-leading benchtop platform.
Expand your capabilities. Achieve 1.8 billion single-end reads per run to power the most data-intensive applications on your benchtop sequencer.
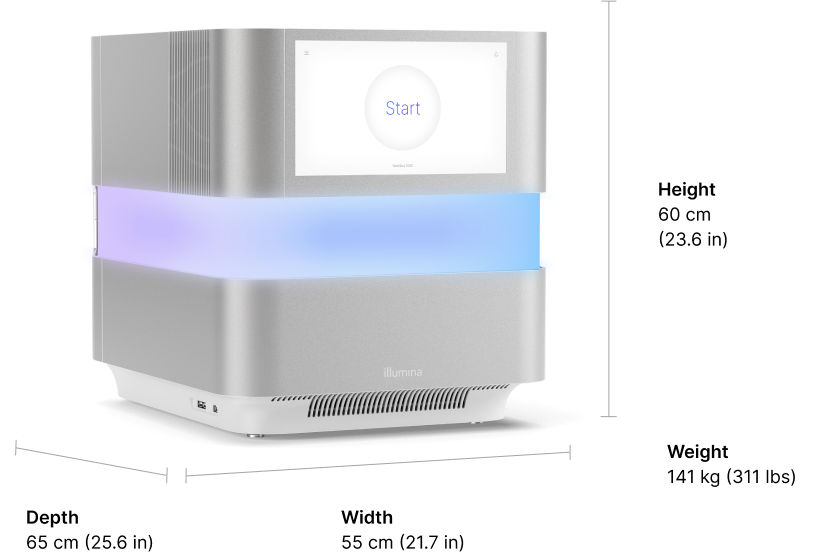


System specifications
a. Specifications based on Illumina PhiX control library at supported cluster densities.
With the help of the NextSeq 2000 and AI, Dyno Therapeutics is powering a new approach to gene therapy. Hear from Eric Kelsic and Ben Noyes about how they are accelerating drug discovery with innovative, scalable technologies.

The NextSeq 1000 and NextSeq 2000 Systems provide a comprehensive workflow that includes user-friendly run setup, a wide ecosystem of compatible library prep kits, load-and-go operation, and integrated, onboard secondary analysis.
Track samples and manage runs easily and efficiently with cloud-based and on-premises options.
Prepare libraries with a wide range of compatible library preparation kits.
Generate sequences with scalable output, efficient run times, and high data quality.
Analyze using onboard DRAGEN secondary analysis or stream to the cloud for additional analysis flexibility.
Access BaseSpace apps, instrument performance data, license management, and long-term storage solutions.
Take the guesswork out of your next workflow. The NGS Workflow Finder provides personalized solution recommendations and resources so you can sequence with confidence.
When you partner with Illumina, you’ll get more than just leading-edge technology–you become part of a community with like-minded goals.
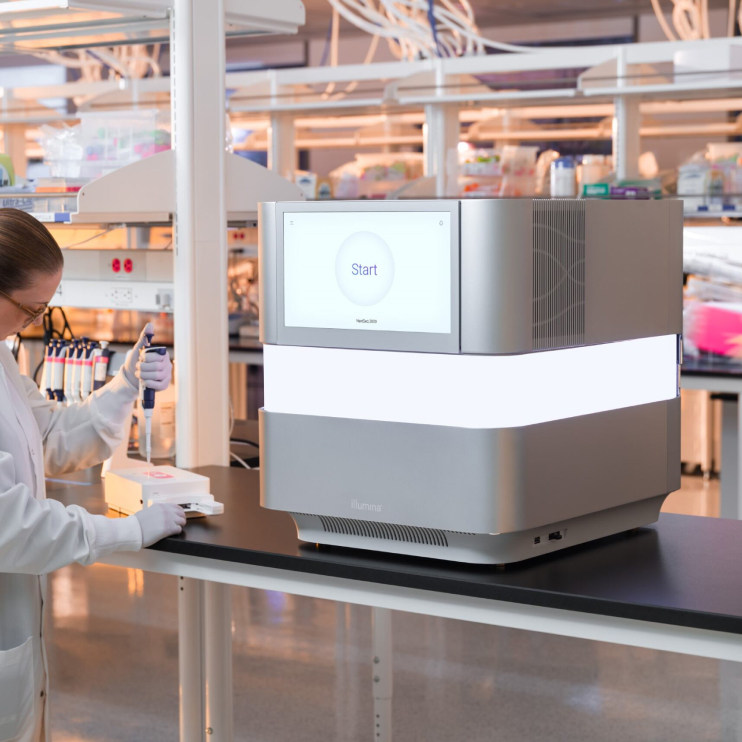

Read how the NextSeq 2000 System provides a more comprehensive and streamlined end-to-end workflow with improved variant calling accuracy and performance compared to the AVITI System from Element Biosciences using real-world data.

Learn about innovations of the NextSeq 1000/2000 workflow, including the use of 3D genomics to study the functional context of GWAS variants, in this webinar.
Perform transformative science with access to multiple emerging omics applications.
|
|
NextSeq 1000 and NextSeq 2000 Systems |
|
|
| Maximum output per runa,b | 15 Gb | 120 Gb | 540 Gbc | 8 Tb |
| Maximum single reads per runb | 25M | 400M | 1.8Bc | 26B |
| Maximum read length | 2 × 300 bp | 2 × 150 bp | 2 × 300 bp | 2 × 150 bp |
| Informatics optionsd | On-premises All DRAGEN Pipelines Cloud DRAGEN Pipelines available on BaseSpace Sequence Hub and Illumina Connected Analytics |
On-premises All DRAGEN Pipelines Cloud DRAGEN Pipelines available on BaseSpace Sequence Hub and Illumina Connected Analytics |
On-instrument Popular DRAGEN Pipelines available onboard On-premises All DRAGEN Pipelines Cloud DRAGEN Pipelines available on BaseSpace Sequence Hub and Illumina Connected Analytics |
On-instrument Popular DRAGEN Pipelines available onboard On-premises All DRAGEN Pipelines Cloud DRAGEN Pipelines available on BaseSpace Sequence Hub and Illumina Connected Analytics |
a. Based on Illumina PhiX control library at supported cluster densities.
b. Data is based on single flow cell output. The NovaSeq X Plus System is capable of single flow cell runs or dual flow cell runs. The NovaSeq X System is capable of single flow cell runs.
c. Specifications are based on a P4 flow cell run. P4 flow cells are available for the NextSeq 2000 System only.
d. View key DRAGEN applications for more details.
Interested in learning more about the flexibility of NextSeq 1000 and NextSeq 2000 Systems? Contact us today to answer your questions.
Your email address is never shared with third parties.