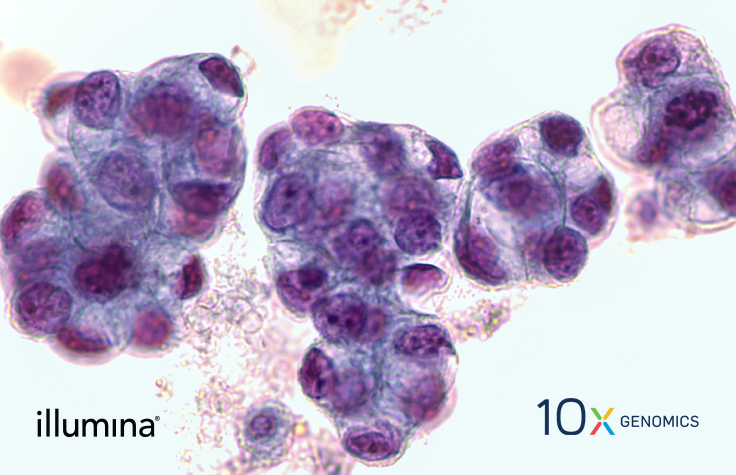
Tumor Microenvironment Analysis
Assessing the Tumor Milieu with NGS
Tumor tissues vary considerably in their microenvironment, and the existence of other cell types can influence the ability of the immune system to infiltrate and attack tumor cells. As promising new therapies evolve, there is an increasing need to identify cancer immunotherapy biomarkers to guide their appropriate application.
NGS can provide careful analysis of the cancer genome, and efficiently assess the tumor microenvironment as a real-time, highly sensitive monitor of immune marker expression in response to tumor growth or treatment. NGS analysis can be used to characterize the immune cell repertoire, identify various cell populations in the microenvironment, and comprehensively quantify gene expression of thousands of targets simultaneously.
Harnessing the Tumor Microenvironment
In this webinar, immuno-oncology leaders explore new approaches for assessing the potential of harnessing the tumor immune microenvironment for therapeutic benefit.
RNA-Seq for Tumor Microenvironment Analysis
RNA analysis may help to identify aspects of the tumor microenvironment that can influence the effectiveness of checkpoint immunotherapies, such as inductive and inhibitory cytokines, and local recruitment of other cell types that can inhibit the T-cell response.
Epigenetic Analysis of Cancer Samples
Studies of epigenetic alterations in cancer, such as aberrant methylation and altered transcription factor binding, can provide insight into important tumorigenic pathways. NGS and microarray technologies can detect altered methylation and other epigenetic patterns in cancer.
Learn More About Cancer EpigeneticsTumor Microenvironment & Immunotherapy Research Articles
Exploring the Tumor Microenvironment
Single-cell sequencing proves invaluable in detecting intracellular communication in tumors.
Read Interview
Immunotherapies Using Tumor-Specific HLA Ligands
RNA-Seq and HLA typing using NGS are powering the Immatics target discovery platform.
Read Interview
Cancer and Immune System Research Review
High-throughput sequencing has shown remarkable utility in cancer and immunology research, as well as in the development of individualized immunotherapy.
Read ReviewFeatured Tumor Microenvironment Analysis Webinars
Characterizing the Tumor Microenvironment in Metastatic Prostate Cancer
A spatial genomics approach to defining the microenvironment landscape of molecular subtypes of metastatic prostate cancer reveals patterns that may predict the utility of immune-based therapies.
View WebinarBuilding a Single Cell and Spatial Tumor Immune Atlas for Precision Oncology
In this webinar, Dr. Holger Heyn of the Barcelona Institute of Science and Technology discusses efforts to combine single cell and spatial transcriptomics to generate a comprehensive tumor immune atlas across cancer types.
View WebinarCancer Research Bioinformatics
Cancer researchers need simple-to-use bioinformatics applications to gain insights from tumor microenvironment studies and other complex genomic data. BaseSpace Correlation Engine allows biologists and researchers to analyze their data in the greater biological context of a highly curated multi-omic datasets with push button analytics requiring no specialized bioinformatics skills. The retrospective analysis of clinical trial data can help inform future cancer research and drug development programs.
Learn more about BaseSpace Correlation Engine.
Informatics Applications for Immuno-oncology Research Using NGS
Improved informatics tools enable tumor microenvironment analysis and neoantigen discovery.
Tumor Microenvironment Analysis Workflow
Illumina offers several library preparation and sequencing options with access to data analysis options for tumor microenvironment analysis. Streamlined workflows and flexible kit configurations accommodate multiple study designs.
Approximately 90% of the world’s sequencing data are generated using Illumina sequencing by synthesis (SBS) chemistry.*
Click on the below to view products for each workflow step.
Illumina DNA Prep with Enrichment uses a fast, user-friendly workflow. On-bead tagmentation chemistry is combined with a simplified, single hybridization protocol to reduce total workflow time.
TruSeq DNA ExomeTruSeq DNA Exome is a cost-effective library preparation and exome enrichment solution.
TruSeq RNA ExomeThe TruSeq RNA Exome library prep kit provides a low-cost solution for analyzing human RNA isolated from limited or low-quality samples, including FFPE.
Transcriptome profiling of hundreds to tens of thousands of single cells in a single experiment.
TruSight Oncology 500Assay targeting multiple variant types, including tumor mutational burden (TMB) and microsatellite instability (MSI), even from low-quality samples.
Flexible configurations that support up to 12 exomes per run.
NextSeq 1000 & 2000 SystemsThese systems are redesigned from the ground up to maximize future proofing, offering sequencing power for high-throughput applications.
Flexibility and unprecedented throughput for virtually any genome, sequencing method, and scale of project.
BaseSpace Sequence Hub offers a wide variety of NGS data analysis apps. Use the RNA-Seq filter to find apps for analyzing RNA sequencing data.
Uses the Isaac Genome Alignment Software and the Isaac Variant Caller for exome data analysis.
Additional Resources
Stratifying Subjects for Immunotherapy Research
Genomics scientists discuss how NGS can help stratify subjects for clinical research.
Interested in receiving newsletters, case studies, and information on cancer genomics?
Sign Up*Data calculations on file, Illumina, Inc, 2015.