
NovaSeq X Series ordering
Advanced chemistry, optics, and informatics combine to deliver exceptional sequencing speed and data quality, outstanding throughput, and scalability.
Prepare whole-transcriptome sequencing libraries from blood-derived RNA with an efficient workflow that removes ribosomal RNA and globin mRNA in a single step.
Assay time
Hands-on time
Input quantity
TruSeq Stranded Total RNA with Ribo-Zero Globin delivers a clear and comprehensive view of the transcriptome from blood-derived total RNA, with fast, efficient sequencing library preparation.
These kits couple the benefits of Ribo-Zero ribosomal RNA reduction chemistry with RNA sequencing (RNA-Seq) technology for whole-transcriptome analysis of human, mouse, or rat samples.
Whole-transcriptome analysis of blood-derived RNA requires the removal of two forms of abundant RNA—ribosomal RNA (both cytoplasmic and mitochondrial) as well as globin mRNA, which is present in high levels in whole blood.
Traditional removal methods require two independent steps. This means the need for additional reagents, a longer workflow, and more input RNA lost. TruSeq Stranded Total RNA with Ribo-Zero Globin leverages Ribo-Zero chemistry to efficiently remove both forms of abundant RNA in a single, rapid step.
| Assay time | 11.5 hr |
|---|---|
| Automation capability | Liquid handling robot(s) |
| Content specifications | Captures coding RNA plus multiple forms of non-coding RNA |
| Description | For whole-transcriptome sequencing studies of blood-derived RNA. Depletes cytoplasmic and mitochondrial rRNA plus globin mRNA. |
| Hands-on time | 5.5 hr |
| Input quantity | 0.1 – 1 ug high-quality purified total RNA from blood |
| Instruments | NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, NovaSeq X System, NextSeq 500 System, NovaSeq 6000 System, NovaSeq X Plus System |
| Mechanism of action | Bead-based rRNA depletion, cDNA synthesis, and PCR |
| Method | Whole-transcriptome sequencing |
| Multiplexing | High-throughput kits: Prepare 96 indexed samples using either combinatorial dual indexes or unique dual indexes, Low-throughput kits: Pool up to 12 samples. Or pool up to 24 samples with sets A and B together |
| Nucleic acid type | RNA |
| Specialized sample types | Blood |
| Species category | Mouse, Human, Rat |
| Species details | rRNA Removal: Depletes human, mouse, and rat rRNA (cytoplasmic and mitochondrial) and globin mRNA |
| Technology | Sequencing |
| Variant class | Single nucleotide polymorphisms (SNPs), Gene fusions, Novel transcripts, Transcript variants |
Illumina now offers modular product ordering to enable flexibility in your workflows. You may order any of the following library prep products with any of the following indexes.
TruSeq Stranded Total RNA with Ribo-Zero Globin
| Instrument | Recommended number of samples | Read length |
|---|---|---|
| NextSeq 550 System | RNA Profiling: 6-20 samples per run (based on 20 million reads per sample) |
2 x 75 bp |
| NovaSeq 6000 System | RNA Profiling (samples per run, dual flow cell): S1: 160, S2: 320, S4: 768. Limited by index combinations (dual). Based on 20 million reads. |
≤ 2 × 100 bp |
Analyze both coding RNA and multiple forms of noncoding RNA for a comprehensive view of the transcriptome.
Find FFPE RNA sequencing solutions that yield high-quality results from degraded FFPE tissues and other low-quality samples.
RNA-Seq uses next-generation sequencing to analyze expression across the transcriptome, enabling scientists to detect known or novel features and quantify RNA.
| TruSeq Stranded Total RNA with Ribo-Zero Globin | Illumina Stranded Total RNA Prep with Ribo-Zero Plus or Ribo-Zero Plus Microbiome | |
|---|---|---|
| Assay time | 11.5 hr | ~7 hr |
| Automation capability | Liquid handling robot(s) | Liquid handling robot(s) |
| Content specifications | Captures coding RNA plus multiple forms of non-coding RNA | Captures coding RNA plus multiple forms of non-coding RNA |
| Description | For whole-transcriptome sequencing studies of blood-derived RNA. Depletes cytoplasmic and mitochondrial rRNA plus globin mRNA. | For the preparation of whole-transcriptome sequencing-ready libraries from low RNA inputs and low-quality samples that supports a wide range of RNA-Seq applications. Includes Ribo-Zero Plus for single-tube depletion of rRNA and globin RNA from multiple species to facilitate rich transcriptome analyses. |
| Hands-on time | 5.5 hr | < 3 hr |
| Input quantity | 0.1 – 1 ug high-quality purified total RNA from blood | 1–1000 ng total RNA for standard-quality RNA samples. 10 ng total RNA minimum recommended for optimal performance and FFPE samples |
| Instruments | NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, NovaSeq X System, NextSeq 500 System, NovaSeq 6000 System, NovaSeq X Plus System | NextSeq 550 System, NextSeq 2000 System, NextSeq 1000 System, NovaSeq X System, NextSeq 500 System, NovaSeq 6000 System, NovaSeq X Plus System |
| Mechanism of action | Bead-based rRNA depletion, cDNA synthesis, and PCR | Enzymatic rRNA depletion, ligation-based addition of adapters and indexes |
| Method | Whole-transcriptome sequencing | Whole-transcriptome sequencing, Metatranscriptome sequencing |
| Multiplexing | High-throughput kits: Prepare 96 indexed samples using either combinatorial dual indexes or unique dual indexes, Low-throughput kits: Pool up to 12 samples. Or pool up to 24 samples with sets A and B together | Up to 384 Unique Dual Indexes (UDIs) |
| Nucleic acid type | RNA | RNA |
| Specialized sample types | Blood | Microbiome samples, Low-input samples, FFPE tissue |
| Species category | Mouse, Human, Rat | Mouse, Human, Rat, Bacteria |
| Species details | rRNA Removal: Depletes human, mouse, and rat rRNA (cytoplasmic and mitochondrial) and globin mRNA | Ribo-Zero Plus depletes abundant transcripts from multiple species including: human cytoplasmic and mitochondrial rRNA, mouse rRNA, rat rRNA, bacteria (E. coli and B. subtilis) rRNA, and human beta globin transcripts Ribo-Zero Plus Microbiome depletes abundant transcripts from multiple species including: those depleted in Ribo-Zero Plus as well as diverse microbial species and ATCC MSA-2002, MSA-2005, and MSA-2006 |
| Technology | Sequencing | Sequencing |
| Variant class | Single nucleotide polymorphisms (SNPs), Gene fusions, Novel transcripts, Transcript variants | Single nucleotide polymorphisms (SNPs), Gene fusions, Novel transcripts, Transcript variants |
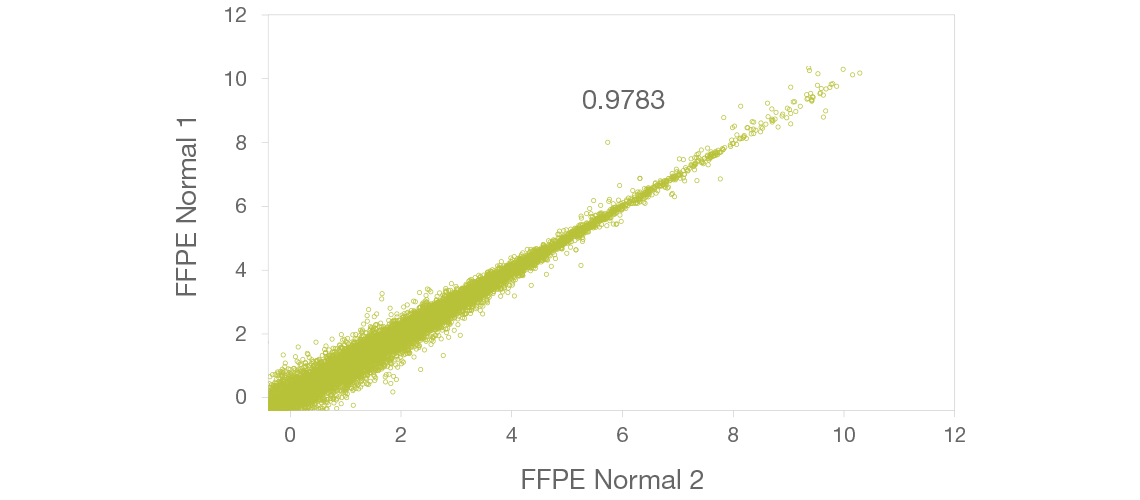

Technical Replicates
Technical replicates of FFPE tissue show high concordance, indicating robust library prep performance. Axes are log2(FPKM). R2 value is shown.
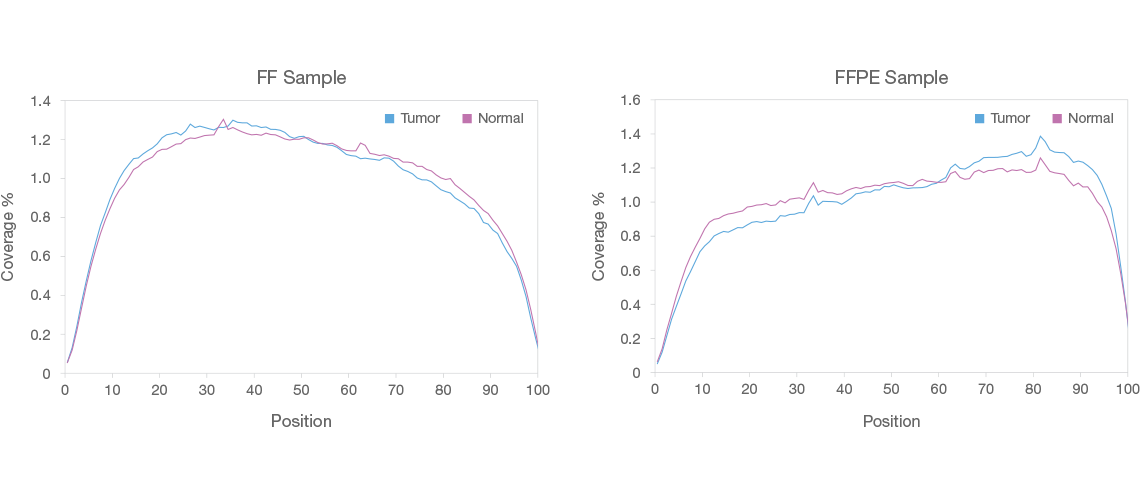

Even Coverage Across Transcripts
TruSeq Stranded Total RNA gives excellent coverage across the top 1,000 expressed transcripts in both fresh-frozen (FF, top) and FFPE (bottom) tumor and matched normal breast tissue, with > 98% aligned stranded reads. X-axis: position along transcript, Y-axis = percent coverage of combined reads.
TruSeq® Stranded Total RNA Library Prep Globin (48 Samples)
20020612
Includes RiboZero beads to deplete rRNA and globin mRNA from blood samples, PCR master mix to transcript RNA into cDNA, and reagents for preparing 48 samples. Purchase index adapters and Purification Beads separately.
List Price:
Discounts:
TruSeq® Stranded Total RNA Library Prep Globin (96 Samples)
20020613
Includes RiboZero beads to deplete rRNA and globin mRNA from blood samples, PCR master mix to transcript RNA into cDNA, and reagents for preparing 48 samples. Purchase index adapters and Purification Beads separately.
List Price:
Discounts:
TruSeq RNA CD Index Plate (96 Indexes, 96 Samples)
20019792
TruSeq Stranded RNA HT Kit 96 Samples Adapter Plate Box
List Price:
Discounts:
IDT for Illumina – TruSeq RNA UD Indexes (24 Indexes, 96 Samples)
20020591
New Unique Dual Indexes (UDIs), compatible with TruSeq Stranded mRNA and TruSeq Stranded Total RNA Library Prep Kits. The UDI kit is provided in a plate format, contains 24 UDIs (x4), sufficient for 96 samples.
List Price:
Discounts:
IDT for Illumina – TruSeq RNA UD Indexes v2 (96 Indexes, 96 Samples)
20040871
Includes 96, 8 bp indexes sufficient for labeling 96 samples and is compatible with TruSeq Stranded mRNA and TruSeq Stranded Total RNA Library Prep Kits. Version 2 contains improvements for 8 of the 192 index oligos in plate 20022371, with no changes to the remaining 184. Purchase library preps and probe panels separately.
List Price:
Discounts:
Showing of
Product
Qty
Unit price
Product
Catalog ID
Quantity
Unit price
Limited Use Label License:
The TruSeq Stranded Total RNA with Ribo-Zero Globin Workflow product and its use are the subject of one or more issued and/or pending U.S. and foreign patent applications owned by Max Planck Gesellschaft, exclusively licensed to New England Biolabs, Inc. and sublicensed to Illumina, Inc. The purchase of this product from Illumina, Inc., its affiliates, or its authorized resellers and distributors conveys to the buyer the non-transferable right to use the purchased amount of the product and components of the product by the buyer (whether the buyer is an academic or for profit entity). The purchase of this product does not convey a license under any claims in the foregoing patents or patent applications direct to producing the product. The buyer cannot sell or otherwise transfer this product or its components to a third party or otherwise use the product for the following COMMERICAL PURPOSES: (I) use of the product or its components in manufacturing; or (2) use of the product or its components for therapeutic or prophylactic purposes in human or animals.
Reach out for information about our products and services, or get answers to questions about our technology.
Your email address is never shared with third parties.